-Search query
-Search result
Showing 1 - 50 of 75 items for (author: kudryashev & m)
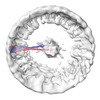
EMDB-18698: 
Structural insights into the activation mechanism of antimicrobial GBP1: Polymeric assembly of GBP1
Method: single particle / : Weismehl M, Chu X, Kutsch M, Lauterjung P, Herrmann C, Kudryashev M, Daumke O

EMDB-18806: 
Structural insights into the activation mechanism of antimicrobial GBP1: Membrane-bound GBP1 oligomer
Method: subtomogram averaging / : Weismehl M, Chu X, Kutsch M, Lauterjung P, Herrmann C, Kudryashev M, Daumke O

PDB-8r1a: 
Model of the membrane-bound GBP1 oligomer
Method: subtomogram averaging / : Weismehl M, Chu X, Kutsch M, Lauterjung P, Herrmann C, Kudryashev M, Daumke O

EMDB-18505: 
Human 80S ribosome structure from pFIB-lamellae reprocessed with TomoBEAR
Method: subtomogram averaging / : Balyschew N, Yushkevich A, Mikirtumov V, Sanchez RM, Sprink T, Kudryashev M

EMDB-17163: 
Structure of the Bacterial Voltage-Gated Sodium Channel NaChBac in Liposomes
Method: subtomogram averaging / : Chang SYS, Kudryashev M

EMDB-17272: 
Structure of RyR1 obtained from SR vesicles tilt series (EMPIAR-10452), reprocessed with TomoBEAR
Method: subtomogram averaging / : Balyschew N, Yushkevich A, Mikirtumov V, Sanchez RM, Sprink T, Kudryashev M

EMDB-17232: 
Purified human apoferritin at 2.8 A resolution from tilt series processed with TomoBEAR
Method: subtomogram averaging / : Balyschew N, Yushkevich A, Mikirtumov V, Sanchez RM, Sprink T, Kudryashev M

EMDB-12191: 
Cryo-EM structure of the Phosphatidylinositol 3-kinase type 2a (PI3KC2a) of the class II PI3K family
Method: single particle / : Lo WT, Zhang Y, Vadas O, Belabed H, Roeske Y, Gulluni F, De Santis MC, Vujicic Zagar A, Stephanowitz H, Hirsch E, Liu F, Daumke O, Nazare M, Kudryashev M, Haucke V

EMDB-13192: 
Structure of knob spiral in P. falciparum-infected erythrocytes
Method: subtomogram averaging / : Chang SYS, Kudryashev M

EMDB-11150: 
Merozoite surface protein 1 (MSP-1) from Plasmodium falciparum, main conformation
Method: single particle / : Dijkman PM, Kudryashev M

EMDB-11151: 
Merozoite surface protein 1 (MSP-1) from Plasmodium falciparum, alternative conformation 2
Method: single particle / : Dijkman PM, Kudryashev M

EMDB-11152: 
Merozoite surface protein 1 (MSP-1) from Plasmodium falciparum, alternative conformation 1
Method: single particle / : Dijkman PM, Kudryashev M

EMDB-11153: 
Merozoite surface protein 1 (MSP-1) from Plasmodium falciparum, alternative conformation 3
Method: single particle / : Dijkman PM, Kudryashev M

EMDB-11154: 
Merozoite surface protein 1 (MSP-1) from Plasmodium falciparum, alternative conformation 4
Method: single particle / : Dijkman PM, Kudryashev M

EMDB-11155: 
Merozoite surface protein 1 (MSP-1) from Plasmodium falciparum, alternative conformation 5
Method: single particle / : Dijkman PM, Kudryashev M

EMDB-11156: 
Plasmodium falciparum merozoite surface protein 1 dimer, conformation 1
Method: single particle / : Dijkman PM, Kudryashev M

EMDB-11157: 
Plasmodium falciparum merozoite surface protein 1 dimer, conformation 2
Method: single particle / : Dijkman PM, Kudryashev M

PDB-6zbc: 
Merozoite surface protein 1 (MSP-1) from Plasmodium falciparum, main conformation
Method: single particle / : Dijkman PM, Kudryashev M

PDB-6zbd: 
Merozoite surface protein 1 (MSP-1) from Plasmodium falciparum, alternative conformation 2
Method: single particle / : Dijkman PM, Kudryashev M

PDB-6zbe: 
Merozoite surface protein 1 (MSP-1) from Plasmodium falciparum, alternative conformation 1
Method: single particle / : Dijkman PM, Kudryashev M

PDB-6zbf: 
Merozoite surface protein 1 (MSP-1) from Plasmodium falciparum, alternative conformation 3
Method: single particle / : Dijkman PM, Kudryashev M

PDB-6zbg: 
Merozoite surface protein 1 (MSP-1) from Plasmodium falciparum, alternative conformation 4
Method: single particle / : Dijkman PM, Kudryashev M

PDB-6zbh: 
Merozoite surface protein 1 (MSP-1) from Plasmodium falciparum, alternative conformation 5
Method: single particle / : Dijkman PM, Kudryashev M

PDB-6zbj: 
Plasmodium falciparum merozoite surface protein 1 dimer, conformation 1
Method: single particle / : Dijkman PM, Kudryashev M

PDB-6zbl: 
Plasmodium falciparum merozoite surface protein 1 dimer, conformation 2
Method: single particle / : Dijkman PM, Kudryashev M

EMDB-11905: 
In situ subtomogram averaging structure of the type III secretion system of yersinia enterocolitica - sorting platform
Method: subtomogram averaging / : Berger C, Ravelli RBG, Lopez-Iglesias C, Kudryashev M, Diepold A, Peters PJ

EMDB-11906: 
In situ subtomogram averaging structure of the type III secretion system of yersinia enterocolitica - basal body
Method: subtomogram averaging / : Berger C, Ravelli RBG, Lopez-Iglesias C, Kudryashev M, Diepold A, Peters PJ

EMDB-11907: 
In situ subtomogram averaging structure of the type III secretion system of yersinia enterocolitica - needle tip
Method: subtomogram averaging / : Berger C, Ravelli RBG, Lopez-Iglesias C, Kudryashev M, Diepold A, Peters PJ

EMDB-10691: 
5-HT3A receptor in Salipro (apo, C5 symmetric)
Method: single particle / : Zhang Y, Dijkman PM, Zou R, Zandl-Lang M, Sanchez RM, Eckhardt-Strelau L, Koefeler H, Vogel H, Yuan S, Kudryashev M

EMDB-10692: 
Serotonin-bound 5-HT3A receptor in Salipro
Method: single particle / : Zhang Y, Dijkman PM, Zou R, Zandl-Lang M, Sanchez RM, Eckhardt-Strelau L, Koefeler H, Vogel H, Yuan S, Kudryashev M

EMDB-10693: 
5-HT3A receptor in Salipro (apo, asymmetric)
Method: single particle / : Zhang Y, Dijkman PM, Zou R, Zandl-Lang M, Sanchez RM, Eckhardt-Strelau L, Koefeler H, Vogel H, Yuan S, Kudryashev M

PDB-6y59: 
5-HT3A receptor in Salipro (apo, C5 symmetric)
Method: single particle / : Zhang Y, Dijkman PM, Zou R, Zandl-Lang M, Sanchez RM, Eckhardt-Strelau L, Koefeler H, Vogel H, Yuan S, Kudryashev M

PDB-6y5a: 
Serotonin-bound 5-HT3A receptor in Salipro
Method: single particle / : Zhang Y, Dijkman PM, Zou R, Zandl-Lang M, Sanchez RM, Eckhardt-Strelau L, Koefeler H, Vogel H, Yuan S, Kudryashev M

PDB-6y5b: 
5-HT3A receptor in Salipro (apo, asymmetric)
Method: single particle / : Zhang Y, Dijkman PM, Zou R, Zandl-Lang M, Sanchez RM, Eckhardt-Strelau L, Koefeler H, Vogel H, Yuan S, Kudryashev M

EMDB-10840: 
Structure of RyR1 in apo/closed state in native membrane
Method: subtomogram averaging / : Chen W, Sanchez R, Zhang Y, Dietrich L, Kudryashev M

EMDB-10834: 
Tobacco mosaic virus (TMV) structure determined by a hybrid STA-SPA workflow
Method: subtomogram averaging / : Sanchez RM, Zhang Y, Chen W, Dietrich L, Kudryashev M

EMDB-10637: 
Structure of ryanodine receptor 1 in apo state in native membrane
Method: subtomogram averaging / : Chen W, Kudryashev M

EMDB-10638: 
Structure of Ryanodine Receptor 1 in the presence of ryanodine in native membrane (with Calcium)
Method: subtomogram averaging / : Chen W, Kudryashev M

EMDB-10639: 
Structure of ryanodine receptor 1 in apo state in native membrane with the T-tubule membrane
Method: subtomogram averaging / : Chen W, Kudryashev M
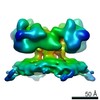
EMDB-10640: 
Rabbit ryanodine receptor 1 in the presence of 1mM Calcium in native membrane
Method: subtomogram averaging / : Chen W, Kudryashev M

EMDB-10641: 
Structure of ryanodine receptor 1 in native membrane in the presence of DDM and EDTA
Method: subtomogram averaging / : Chen W, Kudryashev M

EMDB-10642: 
Structure of rabbit ryanodine receptor 1 in apo state in native membrane (class 1)
Method: subtomogram averaging / : Chen W, Kudryashev M

EMDB-10643: 
Structure of rabbit ryanodine receptor 1 in apo state in native membrane (class 2)
Method: subtomogram averaging / : Chen W, Kudryashev M

EMDB-10644: 
Structure of ryanodine receptor 1 in apo state in native membrane (class 3)
Method: subtomogram averaging / : Chen W, Kudryashev M

EMDB-10645: 
Structure of ryanodine receptor 1 in apo state in native membrane (class 4)
Method: subtomogram averaging / : Chen W, Kudryashev M

EMDB-10646: 
Structure of ryanodine receptor 1 in native membrane in the presence of 5mM Calcium
Method: subtomogram averaging / : Chen W, Kudryashev M

EMDB-4584: 
Structure and assembly of the mitochondrial membrane remodelling GTPase Mgm1
Method: subtomogram averaging / : Faelber K, Dietrich L, Noel J, Wollweber F, Pfitzner A, Muehleip A, Sanchez R, Kudryashev M, Chiaruttin N, Lilie H, Schleger J, Rosenbaum E, Hessenberger M, Matthaeus C, Noe F, Roux A, van der Laan M, Kuehlbrandt W, Daumke O
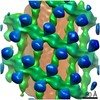
EMDB-10062: 
Structure of s-Mgm1 decorating the outer surface of tubulated lipid membranes
Method: subtomogram averaging / : Faelber K, Dietrich L, Noel JK, Sanchez R, Kudryashev M, Kuehlbrandt W, Daumke O
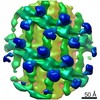
EMDB-10063: 
Structure of s-Mgm1 decorating the outer surface of tubulated lipid membranes in the GTPgammaS bound state
Method: subtomogram averaging / : Faelber K, Dietrich L, Noel JK, Sanchez R, Kudryashev M, Kuelbrandt W, Daumke O

EMDB-10064: 
Structure of s-Mgm1 decorating the inner surface of tubulated lipid membranes
Method: subtomogram averaging / : Faelber K, Dietrich L, Noel JK, Sanchez R, Kudryashev M, Kuelbrandt W, Daumke O
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
